Calibration refers to improving our model such that predicted probability distribution is similar to the probability observed in training data.
Calibration plot (=q-q plot)
Assume we are predicting probabilities for 2 classes .
To produce a calibration plot
- sort by predicted probability
- Define bins between and and compute
- plot against expected true probability for the bin .
How do we derive ? Think of it as first sampling a probability , then, if sample a random variable: taking value in .
What is the fraction of positives (i.e. the expectation of ) as varies between and ?
Perfect calibration plot should be identity:
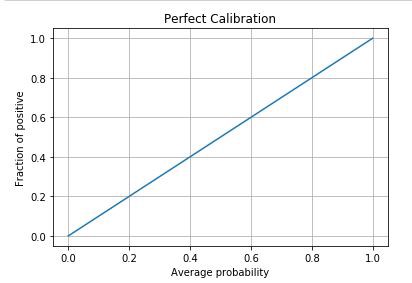
Pseudo-code:
import math
import seaborn as sns
def plot_calibration_curve_binary(label, y_proba):
# expected true proba within a bin.
def expected_proba(l, r):
return 2/3 * (r**3 - l**3) / (r**2 - l**2)
df = pd.DataFrame({
"label": label,
"y_proba": y_proba,
})
# rule of thumb for number of bars N given number of data points n: N = ceil(log2(n) + 1)
N = math.ceil(math.log2(len(df) + 1))
bins = pd.cut(df["y_proba"], bins=np.linspace(0, 1, N))
calibration_df = df.groupby(bins, observed=False).agg(mean=("label", "mean"), count=("label", "count"))
calibration_df["expected_proba"] = list(map(lambda x: expected_proba(x.left, x.right), calibration_df.index))
sns.scatterplot(x=calibration_df["expected_proba"], y=calibration_df["mean"], label="y_proba")
sns.lineplot(x=[0, 1], y=[0, 1], color="red") # perfect calibration line
sns.despine() # remove top and right spines
Sigmoid / Platt calibration
Logistic regression on our model output:
Optimized over and .
Isotonic regression
Let . Isotonic regressions seeks weighted least-squares fit s.t. whenever .
Objective is: assuming the 's are ordered.
This yields a piecewise constant non-decreasing function. To solve this we use the pool adjacent violators algorithm. See these notes.